Nucleic Acid Chemistry
Chemistry of Oxidative DNA Damage & Repair
(supported by NIH: R01 CA090689)
Cells thrive on redox processes, but when free radical oxygen species get out of control, the DNA base guanine (G) is a target. Complex heterocyclic chemistry ensues, and the formation of various oxidized lesions in DNA depends upon the intricacies of cellular chemistry as well as the DNA sequence and secondary structure. G oxidation to 8-oxoG (OG) is somewhat mutagenic if not repaired, and an accumulation of DNA damage and mutations can lead to cancer.

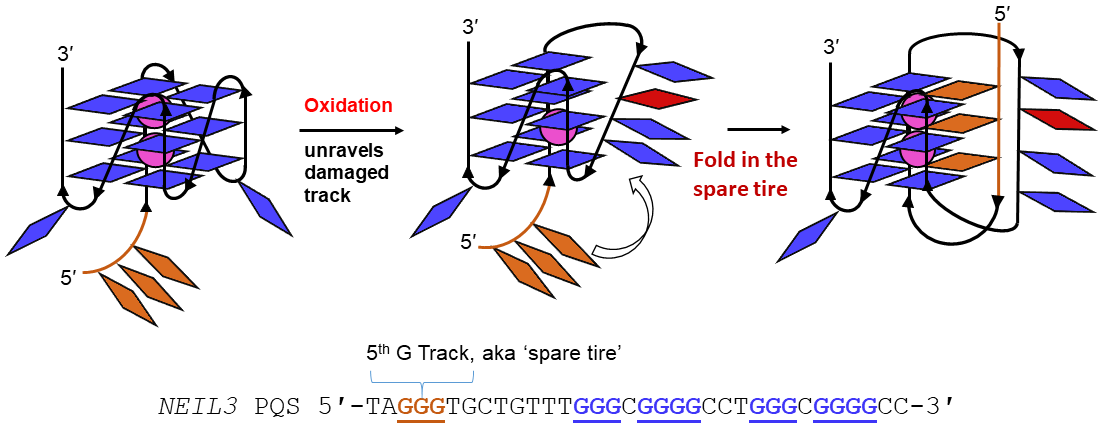
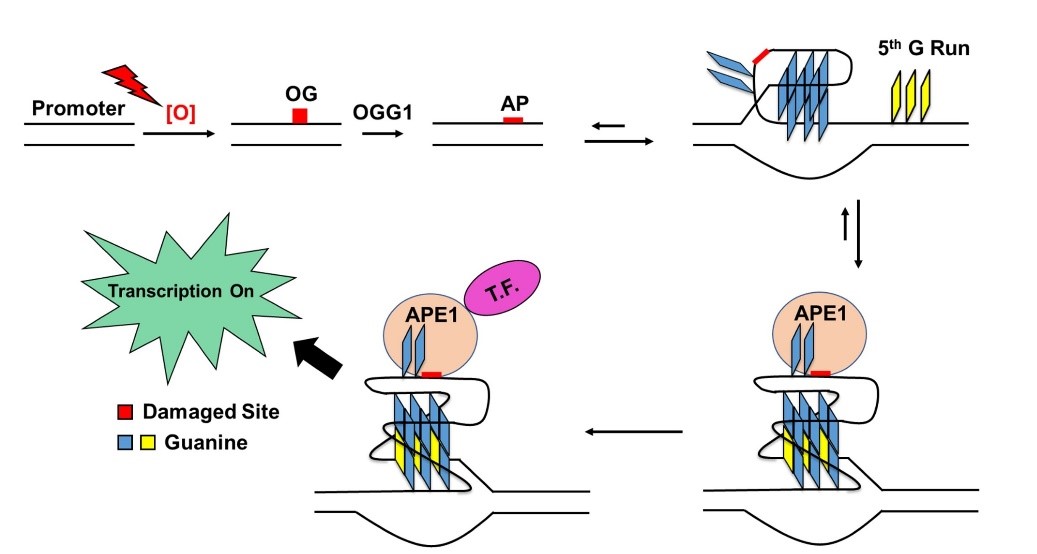
On the flip side, we have shown that OG in a G-quadruplex (G4) sequence in a gene promoter can be a natural response to oxidative stress by turning on (or off) gene expression via base excision repair. In this sense, what was thought to be ‘damage’ might actually be called ‘epigenetic.’ We are examining the molecular mechanisms of this pathway.
Sequencing for Base Modifications in DNA
(supported by NIH: R01 GM129267 & R01 GM093099)
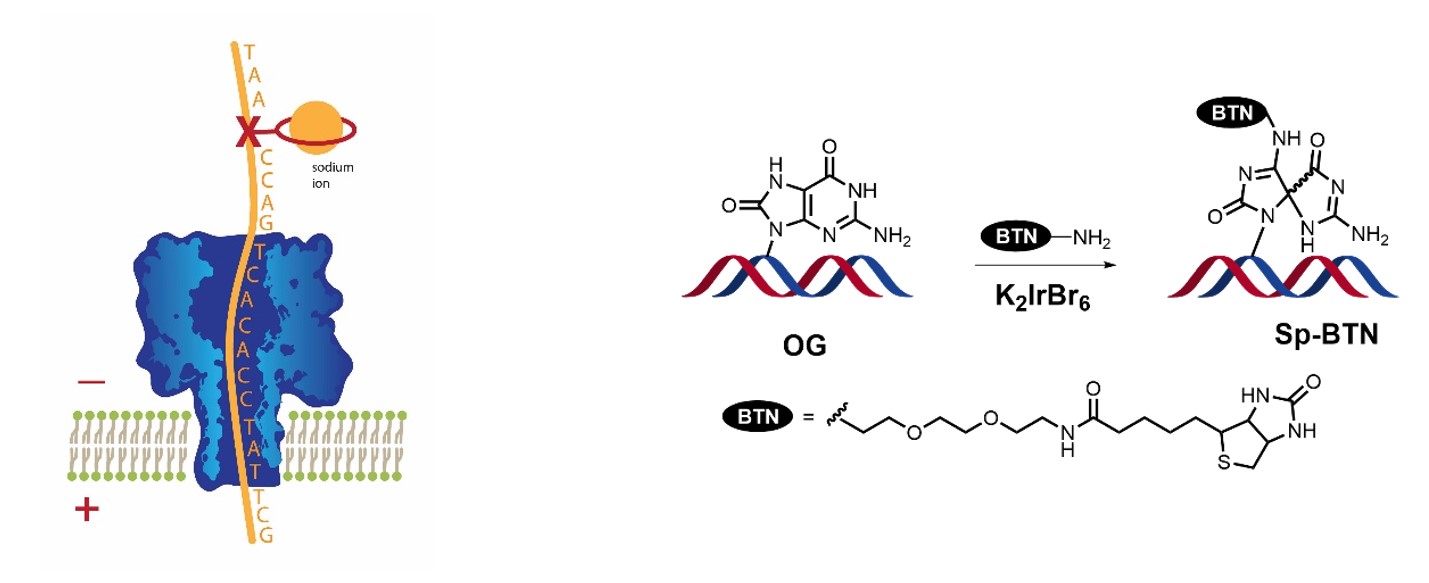
A key tenet of our oxidative ‘damage/epigenetics’ hypothesis is that guanine oxidation is focused in specific regions of the genome, notably G-rich regions such as telomeres, gene promoters, and other regulatory elements. We continue to investigate approaches to sequencing DNA for base modifications such as OG, Sp and Gh. Here’s a timeline:
- 2011, 2012 – Single-molecule analysis of AP and OG using the α-hemolysin nanopore
- 2015 – Counting OG in telomeric sequences via the nanopore
- 2015 – Using BER glycosylases to locate OG, AP and hydantoins with unnatural base pair amplification
- 2016 – The gap ligation strategy for Sanger sequencing for OG, AP and other lesions
- 2017 – OG-Seq: Whole genome Next-gen sequencing after chemical pull-down
- 2020 and beyond – Taking OG-seq to the next level with unbiased sequencing at higher resolution
Base Modifications in RNA
(supported by NIH: R01 GM093099)
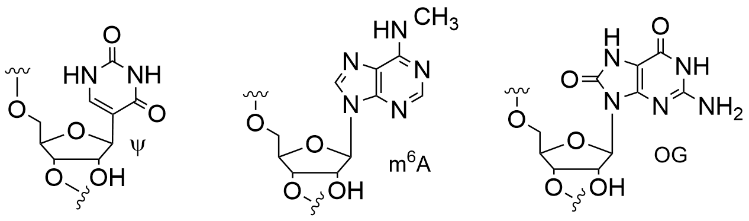
As in DNA, base modifications in RNA play key roles but they are rare compared to unmodified bases. Why are certain bases in specific locations subject to installation of modifications? Both sequence and secondary structure can play a role. We found that G-rich sequences in mRNA and viral RNA (Zika, HIV, HepB…) contain potential G-quadruplex sequences, and these co-locate with the epitranscriptomic modification m6A. For other base modifications in RNA, the correlation to folded motifs is less obvious. In collaboration with the Cairns laboratory, we have found methods to sequence for pseudouridine, a modified base that may be correlated to cellular stress.
Quadruplex Nucleic Acids
(supported by NSF CHE1808745 and NIH R01 CA090689)
Beyond the Watson-Crick canonical duplex, most genomes include sequences capable of alternative folding to G-quadruplexes and i-motifs. We are interested in the following questions:
- Which organisms, from mammals to microbes, utilize these sequences to punctuate their genomes?
- How and when to they fold to impact cellular responses to stress including endogenous factors as well as inflammation, photochemical stress, and radiation damage?
- How does sequence impact the folding dynamics? These studies include our ‘spare tire’ hypothesis, and in the i-motif fold, the dependence on the loop composition.