Visualizing Molecules with gOpenMol

gOpenMol is a molecular visualization program written by Leif Laaksonen and available from him at www.csc.fi/gopenmol/ . Versions are available for Windows, Linux, and some Unix platforms.
This tutorial was written by Scott Anderson and applies to both the Win 95/98/NT..., Linux, and UNIX versions of gOpenMol. The tutorial is not intended to be a manual, but rather a simple walk-through showing how to accomplish some of the more common types of visualization. The examples are all from Gaussian 98, however, gOpenMol can be used with a very large variety of quantum chemistry/molecular mechanics programs. The tutorial covers:
1. Installing and running gOpenMol on Windows and some Linux hints
2. Importing and viewing calculated molecular structures in various ways
3. Visualizing molecular orbitals, electron densities, spin densities, etc. in several
different ways
4. Viewing Vibrations or other trajectory information:
A. Animations
B. Vibrations as static vectors
C. Analyzing Trajectories
5. Making movies
6. Selecting and manipulating groups of atoms, measuring molecules, merging structures
and exporting structures
7. Global v.s. Local transformations
8. Automating with scripts
9. Features for Biomolecule Visualization
10. Creating VRML files (3D images and trajectories that can be manipulated over the
web)
11. Customization and other stuff I thought was useful/cool
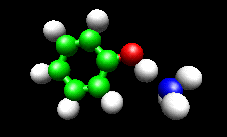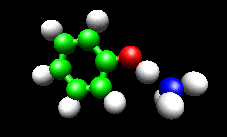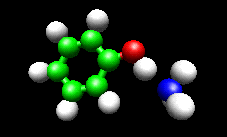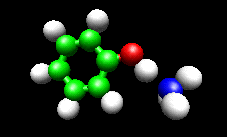
The first example image shows the shape of the total electron density for a molecule thought to be an intermediate in decomposition of methyl cubane. The second image shows a vibration of the phenol cation - ammonia complex that is found experimentally to substantially enhance the cross section for H atom exchange in reaction of state-selected PhOH cation with ammonia.
Download Tutorial.zip The zip file contains sample files used in the tutorial, however, you can just as easily substitute your own files.
Download just the stand-alone xvibs utility
Return to Scott Anderson's home page
